Once identified the best enzymes, it’s time to study them in their context: the carbon positive CO2 fixation pathway. “We want to make sure the enzyme can work well as a team. It’s like in football, you might have great individual players but to score you need a good teamwork, too. Sometimes it makes sense to take out one of the best players and substitute with another option because it interacts better with the rest of the team” explains Tobias Erb (MPI-MP).
To test the interplay of the individual enzymes, the CO2 fixation pathway is first recreated in vitro: a small vessel containing all the ingredients necessary to the pathway i.e. the individual enzymes, the chemical energy (ATP) and its reducing equivalents (NADPH), other cofactors that help the enzyme to exert its function, and a buffer that mimics the cell environment.
The pathway performance is then quantified by measuring the variations in the concentration of the ingredients. This is done by using highly sensitive mass spectrometers that can accurately quantify the amounts of substrates, intermediates and other metabolites for all the reactions that are happening in the vessel.
The team of FutureAgriculture has already tested in vitro the effect of the newly designed pathway onto the CO2 fixing-machinery of the plant. The outcome was extremely encouraging as the pathway showed a beneficial effect on the plant metabolism. This is the very first proof of principle that the pathway is able to improve plant metabolism in the lab condition, and that the pathway if transplanted in vivo has the potential to positively affect plant growth.


 Our partner, Tobias Erb from the Max Planck Institute for Terrestrial Microbiology have been awarded the prestigious
Our partner, Tobias Erb from the Max Planck Institute for Terrestrial Microbiology have been awarded the prestigious 
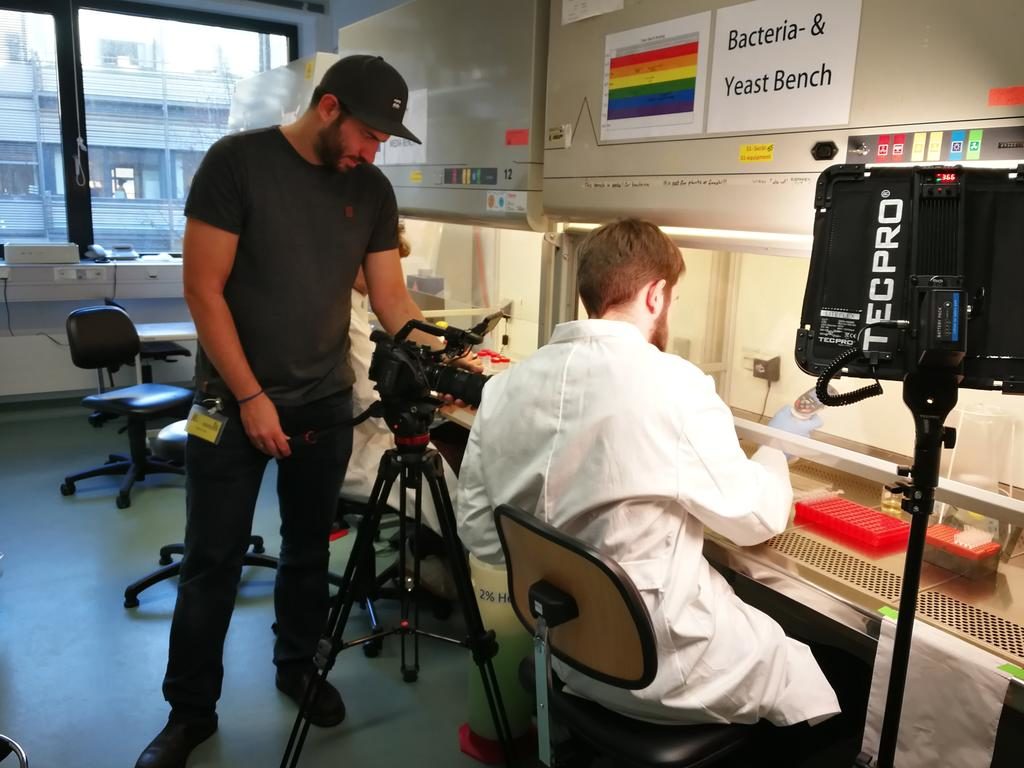 We are currently collaborating with MedioMix to create the official video for FutureAgriculture! The first shooting day was at the MPI-MP in Potsdam – it has been a long day among laboratories, greenhouses and finding the right location for interviews. The official video for FutureAgriculture will be released soon, in the meantime, you can check our other videos on the dedicated
We are currently collaborating with MedioMix to create the official video for FutureAgriculture! The first shooting day was at the MPI-MP in Potsdam – it has been a long day among laboratories, greenhouses and finding the right location for interviews. The official video for FutureAgriculture will be released soon, in the meantime, you can check our other videos on the dedicated  Unleashing the technical present / Nov 30 – Dec 02. Researchers & artists met for this experimental conference in order to explore the meaning and implications of the technosphere. Arren Bar-Even was invited to the SEEDS session – watch the full performance
Unleashing the technical present / Nov 30 – Dec 02. Researchers & artists met for this experimental conference in order to explore the meaning and implications of the technosphere. Arren Bar-Even was invited to the SEEDS session – watch the full performance  Our project coordinator got the chance to interact with artists and scientists on the topic “ESTHETICS get SYNTHETIC: Knowledge Link Through Art & Science” during the workshop organized by KLAS at the
Our project coordinator got the chance to interact with artists and scientists on the topic “ESTHETICS get SYNTHETIC: Knowledge Link Through Art & Science” during the workshop organized by KLAS at the  The 1st Synthetic Biology conference “Made in Germany”, GASB I, and the founding event of the German Association for Synthetic Biology, GASB took place on the 24th and 25th of November 2017 in Marburg, Germany. Several members of the FutureAgriculture team were there: Marieke Scheffen and Jan Zarzycki from MPI-TM and, among the keynote speakers, our project coordinator Arren Bar-Even.
The 1st Synthetic Biology conference “Made in Germany”, GASB I, and the founding event of the German Association for Synthetic Biology, GASB took place on the 24th and 25th of November 2017 in Marburg, Germany. Several members of the FutureAgriculture team were there: Marieke Scheffen and Jan Zarzycki from MPI-TM and, among the keynote speakers, our project coordinator Arren Bar-Even. One of our partners, the Max Plack Institute for Molecular Plant Physiology – MPI-MP, has been selected among the 4 finalists for the European Innovation Radar prize 2017 under the category Excellent Science. This category selects the best cutting-edge science underpinning tomorrow's technological advances. Thanks to this initiative, the FutureAgriculture's team got the chance to pitch their plans for going to market with their EU-funded tech to a jury of experts at the ICT Proposers' Day in Budapest (9 November 2017). More info
One of our partners, the Max Plack Institute for Molecular Plant Physiology – MPI-MP, has been selected among the 4 finalists for the European Innovation Radar prize 2017 under the category Excellent Science. This category selects the best cutting-edge science underpinning tomorrow's technological advances. Thanks to this initiative, the FutureAgriculture's team got the chance to pitch their plans for going to market with their EU-funded tech to a jury of experts at the ICT Proposers' Day in Budapest (9 November 2017). More info